 |
Centre for Australian National Biodiversity
Research
|
The Orchidaceae is the most speciose flowering plant family in the world, containing approximately 30,000 species (Batty et al, 2002). Such high taxonomic diversity is partly the result of the production of large numbers of dust-sized seeds that favours the expression of genetic variability (Batty et al, 2002). This variability has led to a range of highly specific pollination systems within the Orchidaceae, which has in turn resulted in a diverse range of attractive flower morphologies that are often of considerable commercial and cultural interest (Jones and Clements, 2002).
In addition, many orchid species are endangered or threatened as a consequence of direct and indirect anthropogenic factors. These include habitat destruction or alteration and the removal of species from their natural environment for the horticulture trade or amateur collections. Together with the shear diversity of the family, this has resulted in a relatively high number of endangered species when compared with other families (Batty et al, 2002). Relatively little is known about the ecology of the Orchidaceae, and a greater understanding is required in order to provide more information for the conservation of the endangered species at present and in the future.
All members of Orchidaceae require symbiotic interactions with both their pollinators and mycorrhizal fungi in order to germinate and complete their life cycle, and that the availability of the fungal symbionts may play a vital role in determining orchid distribution and diversity (Curtis, 1939; Arditti et al, 1990; Rasmussen, 1995; Otero et al, 2002; Kristiansen et al, 2004; Otero et al, 2004). This renders the Orchidaceae as an extremely complex family and as such, further research on these associations will be important for cultivation and conservation efforts (Bayman et al, 2003).
The Pterostylidinae are a group of Australian terrestrial orchids comprising approximately 250 species historically treated as the polymorphic genus Pterostylis. In a phylogenetic reassessment, this large genus was divided into 16 genera based on morphological data and DNA sequence of the ITS ribosomal region (Jones and Clements, 2002). These genera are: Bunochilus, Crangonorchis, Diplodium, Eremorchis, Hymenochilus, Linguella, Oligochaetochilus, Petrorchis, Pharochilum, Plumatichilos, Pterostylis, Ranorchis, Speculantha, Taurantha, and Urochilus. Recently the diversity of their mycorrhizal fungi is being studied showing that the group is preferment associated to particular clades of Ceratobasidium (Otero & Clements unpublished data).
The aims of this project were three-fold. Firstly, to determine the fungal specificity of the Speculantha genus (sub-tribe Pterostylis) with regards to mycorrhizal fungi from different phylogenetic classification, as well as from different locations. Secondly, a new technique for identifying Pterostylis mycorrhizal fungi based on their DNA fragment lengths of the small ribosomal subunit of the mitochondria was developed. Lastly, to corroborate an existing phylogenetic tree of the nuclear ribosomal internal transcribed spacer (ITS) DNA region with the information from the small ribosomal subunit of the mitochondria. This would give major insights into the evolution of orchid mycorrhizal fungi in Australia and provide important information for aiding the conservation of many orchid species, especially those belonging to the Pterostylis sub-tribe.
The seed germination experiment was designed to use a single batch of Speculantha seeds from one fruit of one individual (avoiding genetic factors in the variation) to investigate their germination with mycorrhizal fungi in parallel symbiotic and asymbiotic experiments.
Four replicates of each of six fungal species treatments were used in this experiment, resulting in the use of 24 divided petri dishes in total. The plates contained cellulose media as plants cannot obtain nutrients from this media alone, and any germination observed can be explained as a result of the presence of the fungi. The fungi used were from four different ITS clades as outlined by an existing ITS phylogenetic tree, and were isolated from hosts located in either eastern Australia (NSW or TAS) or Western Australia.
The Speculantha seeds were surface sterilised prior to their transferal onto one side of the divided plates. Sections of pure live fungal cultures were then transferred to the opposite side of the plates and they were sealed and left in the dark for three weeks. The numbers of seeds in each of the 6 protocorm growth stages of development were then counted for each treatment and the results analysed using a one-way Analysis of Variance (ANOVA).
It has been found in field experiments that the testa may rupture (stage 2 of protocorm development (Fig. 1)) several weeks before fungal infection and before considerable growth takes place, and also that a stage 2 seed may not continue its development into an adult plant. As such, those seeds that had reached stage 3 (Fig.1) or above were deemed to be germinated for the purposes of this experiment. However, this does not give an indication of the overall success of each of the treatments and as such the Growth Index was used. The growth index is a standardised number that weights each stage of development according to its progression toward becoming an adult plant in such a way that the later stages of development are given higher weights (following Otero et al, 2004a and 2004b).
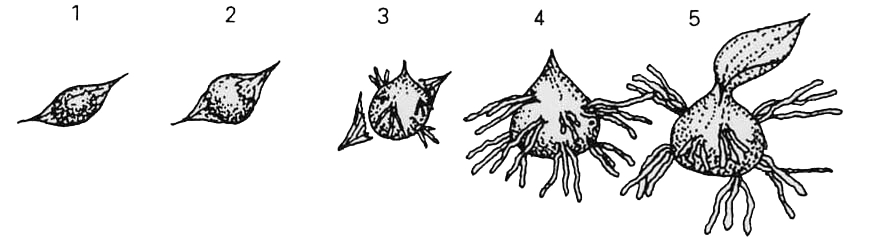
Figure 1: Stages of protocorm development for an orchid seed.
The 34 fungal samples chosen for this project were the same samples used to generate the existing ITS phylogenetic tree that was to be assessed. Thus, the DNA was already extracted and the samples were ready to be used in a Polymerase Chain Reaction (PCR).
The 50m L PCR mix used for each sample contained: 1m L DNA template, 5 m L PCR buffer, 1.25m L (10mM) of each of the primers MLIN3 and ML6, 33.75m L sterile ddH2O, 1m L dNTP (10mM) mix, 7.5m L MgCl2 and 0.5m L Taq polymerase. The PCR products were cleaned using Qiaquick columns (Qiagen Inc., Valencia, CA, USA), and sequenced in both directions using ABI PRISM BigDye Terminator Cycle Sequencing Ready Reaction Mix (Applied Biosystems, Foster City, CA, USA). The sequencing product was precipitated in 60% isopropanol, and sequenced on a ABI 3730 capillary automated DNA sequencer (PE Applied Biosystems, Foster City, CA, USA) at the Australian Genome Research Facility (AGRF, Melbourne, Australia). Both strands of the sequences were edited using the Sequencher 4.0 program (Gene Codes Corp., Ann Arbor, MI, USA), and the resulting consensus sequences were aligned with sequences in GenBank (Table 2) using the ClustalW procedure in Bioedit 5.0.9 (Hall 1999).
The mean percentage of seeds that germinated was found to be significantly different between the fungal treatments and the control (F6, 21,a =0.05= 86.7275, P<0.0001).
It was found that the mean (± SD) growth indices of the fungal treatments were significantly higher than that of the control (F6, 21, a =0.05=82.8492, P<0.0001) (Fig.2) meaning that a greater proportion of the seeds in these treatments had progressed to the later stages of protocorm development.
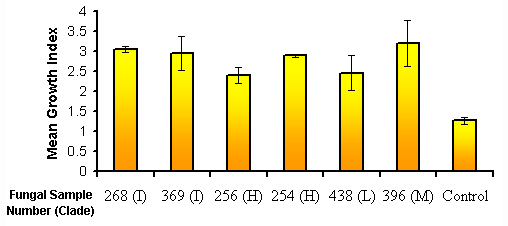
Figure 2: The mean growth index of each of the fungal sample treatments. Error bars are ± SD. Cladogram below graph shows that evolutionary relationship of the samples to one another. Letters in parentheses are the ITS clade of the fungi (Otero & Clements unpublished data).
There was also significant variation in growth index within the fungal treatments themselves. The mean (± SD) growth indices of fungal treatments 268, 369, 396, and 254 (clades I, I, M and H respectively; after Otero and Clements, unpublished data) were significantly higher than that of 256 (clade H) and 438 (clade L) and were not significantly different to each other. Fungal treatments 256 and 438 were not significantly different to each other.
Clade M produced the most successful germination of seeds however this was unexpected as Speculantha mainly associates with fungi from clade I and clade M is the least related clade to clade I (Fig.2). Clade I and clade H followed clade M in germination success, which was expected due to the close relationship of clade H to clade I (Fig 2).
No significant difference was found in the mean growth indices of the seeds germinating with eastern Australian fungi or with western Australian fungi (F1, 22,a =0.05=1.2274, p=0.2799) (Fig 3). This result was unexpected since the distribution of Speculantha is restricted to Eastern Australia. This suggests that it is not the distribution of compatible fungi that is limiting the distribution of Speculantha to eastern Australia; rather it is another factor most likely related to the soil or local environment, for example the existence of suitable habitat that both the orchid and fungi can survive in (Curtis, 1939). It also shows that Speculantha does not have a high specificity for eastern Australian fungi, which may be important for conservation in the future.
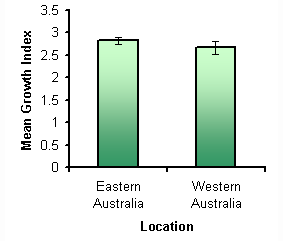
Figure 3: The mean growth indices of the fungal samples isolated from hosts in either eastern Australia (NSW or TAS) or Western Australia. Error bars are ± SD.
Using molecular techniques, it was found that fungi from some of the ITS clades produced MLIN3/ML6 region rDNA fragments of approximately equal length (fig 4). Figure 4 shows that samples from ITS clade M produced approximately equal MLIN3/ML6 region rDNA fragment lengths and that these are longer than those of clades E, T and G, which also produced fragments of approximately equal length.
This suggests that these fungi may actually be identified based solely on their MLIN3/ML6 rDNA fragment length. This would enable the fast and effective identification of the fungi without the need of sequencers, which would also decrease the costs associated with DNA sequencing. These preliminary results suggest further development of this technique needs to be performed in order to assess its ability to identify fungi from clades not tested in this project, and also to improve its accuracy and reliability for future use.
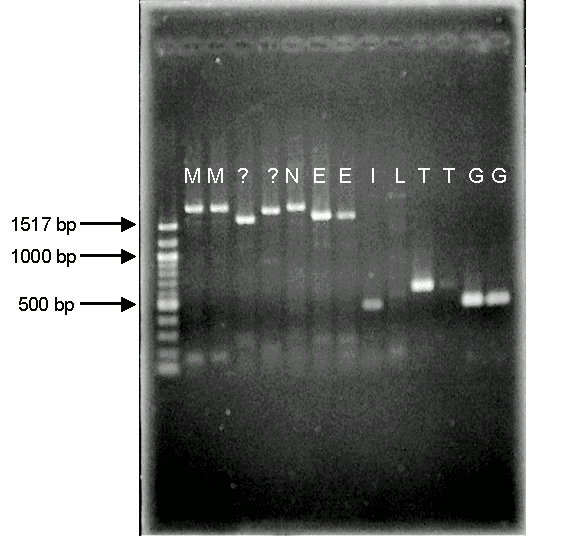
Figure 4: A photograph of an agarose gel stained with ethidium bromide to highlight DNA fragments. DNA samples from ITS clades M, E, T and G show fragments of equal length and these lengths differ between clades. DNA ladder also present showing lengths of DNA bands.
The molecular experiment also revealed that the MLIN3/ML6 region produces fragments that are very large (mostly over 1500 base pairs) and also highly variable. The 34 fungal samples tested produced eight previously unidentified intron types in the MLIN3/ML6 region. Consequently, the ML and ITS rDNA regions could not be directly compared via phylogenetic trees as the ML samples were too variable to be aligned and therefore a phylogenetic tree was unable to be constructed. However, the eight identified intron types were found to correspond to different ITS clades, with each clade containing only one ML intron type (Fig 5). The exception to this is ITS clade E, which contained three ML intron types, however these are specific to clade E. It is recommended that further research be conducted on the eight newly identified introns in the MLIN3/ML6 in order to design primers to target specific introns and also to allow the construction of the MLIN3/ML6 region phylogenetic tree showing its evolution for more thorough comparison with the ITS region.
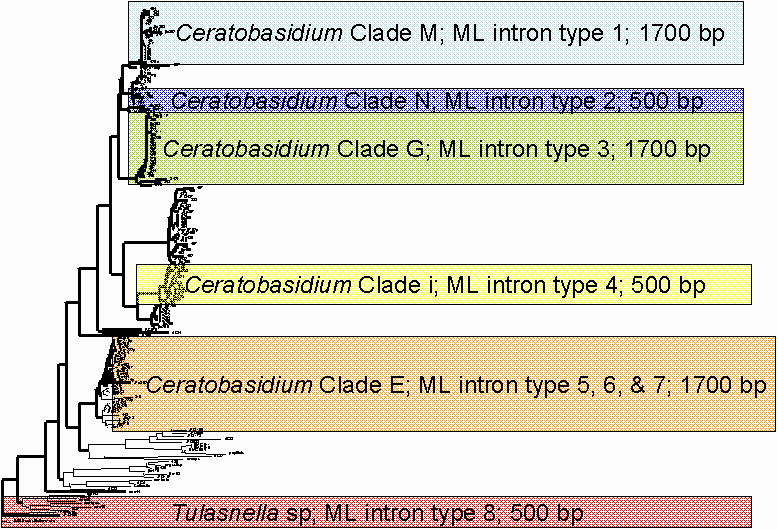
Figure 5: A scaled-down version of the existing ITS phylogenetic tree used for comparison with the ML rDNA region showing the location of each of the ITS clades studied and the type of fungi found in these clades, as well as the location of the eight newly identified ML region intron types and their approximate lengths in base pairs (bp).
Thank you to my supervisors Dr Tupac Otero and visiting academic Dr Paul Bayman for all of their enthusiasm, encouragement, support and advice over the last ten weeks, it is most appreciated. I would also like to thank the staff in the Australian National Herbarium, especially those from the molecular systematics lab for all of their technical help and support. Finally, thank you to the Gatsby Charitable Foundation for economic support to the project, CSIRO, CPBR and the Australian Pastoral Research Trust for giving me the opportunity to participate in this fantastic and rewarding experience.
Arditti J., Ernst R., Yam T.W., Glabe C. (1990) The contributions of orchid mycorrhizal fungi to seed germination: a speculative review. Lindleyana, 5, 249-255
Bayman P., Otero J.T., Ackerman J.D. (2003) Hidden transactions: The curious relationships between orchids and fungi. Orchids. www.orchidweb.org
Batty, A.L., Dixon, K.W., Brundrett, M.C., and Sivasithamparam, K. (2002) Orchid conservation and mycorrhizal associations. In: Microorganisms in Plant Conservation and Biodiversity (Sivasithamparam, K., Dixon, K.W. and Barrett, R.L., eds). Kluwer Academic Publishers pp.195-226
Curtis, J.T. (1939) The relation of specificity of orchid mycorrhizal fungi to the problem of symbiosis. American Journal of Botany, 26, 390-399
Jones D.L., Clements M.A. (2002). A reassessment of Pterostylis R.Br (Orchidaceae). Australian Orchid Research 4: 3-63.
Kristiansen K.A., Taylor D.L., Kjoller R., Rasmussen H.N., Rosendahl S. (2001) Identification of mycorrhizal fungi from single pelotons of Darylorhiza majalis (Orchidaceae) using single-strand conformation polymorphism and mitochondrial ribosomal large subunit DNA sequences. Molecular Ecology, 10, 2089-2093.
Otero, J.T., Ackerman, J.D. and Bayman, P. (2002) Diversity and host specificity of endophytic Rhizoctonia-like fungi from tropical orchids. American Journal of Botany, 89(11), 1852-1858
Otero, J.T., Ackerman, J.D., and Bayman, P. (2004a) Differences in mycorrhizal preferences between two tropical orchids. Molecular Ecology, 13(8), 2393-2404
Otero, J.T., Ackerman, J.D. and Bayman, P. (2004b) Variation in mycorrhizal performance in the epiphytic orchid Tolumnia variegata in vitro: the potential for natural selection. Evolutionary Ecology, 00, 1-15, in press.
Rasmussen, H.N. (1995) Terrestrial orchids: From Seeds to Mycotrophic Plants. Cambridge University Press, New York.